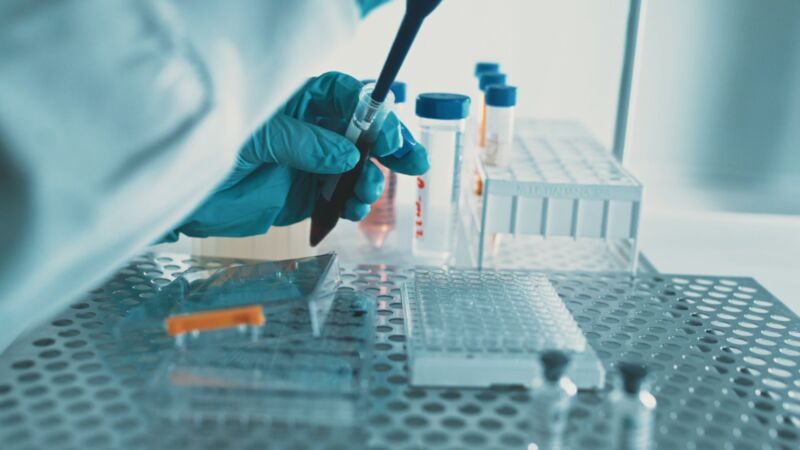
Your metabolomics service experts
We unlock the potential of your research by using the power of metabolomics
- Standard lead time of 4-6 weeks
- Collaborate directly with metabolomics experts
- Versatile methods optimized to your needs
- Cutting edge LC-MS/MS instruments
Applications
MS-Omics has experience with a wide range of applications
Metabolomics methods
MS-Omics offers services and collaborations to assist you with analytical chemistry and advanced data handling